胰腺癌发生发展的分子机制与靶向治疗策略
DOI: 10.12449/JCH260234
-
摘要: 胰腺癌恶性程度高,5年生存率不足10%,其分子机制以Kirsten大鼠肉瘤病毒癌基因同源物(KRAS)突变(90%)、肿瘤蛋白p53/细胞周期蛋白依赖性激酶抑制因子2A/SMAD家族成员4失活及表观遗传异常(DNA甲基化、非编码RNA等)为核心,驱动肿瘤进展。近年来,靶向治疗取得突破,如KRAS原癌基因甘氨酸12→半胱氨酸突变抑制剂(Sotorasib、Adagrasib)和非共价KRAS原癌基因甘氨酸12→半胱氨酸突变抑制剂的临床应用;针对表皮生长因子受体、DNA修复(多腺苷二磷酸核糖聚合酶抑制剂)及免疫微环境(靶向程序性细胞死亡受体1及其配体联合疗法)的策略也显著提升疗效。然而,耐药性与肿瘤异质性仍是挑战。未来需结合精准医疗和联合疗法以改善预后。Abstract: Pancreatic cancer (PC) has a high degree of malignancy and a 5-year survival rate of <10%, with the core molecular mechanisms of Kirsten rat sarcoma viral oncogene homolog (KRAS) mutations (90%), inactivation of tumor protein p53/cyclin-dependent kinase inhibitor 2A/SMAD family member 4, and epigenetic dysregulation (including DNA methylation and non-coding RNA alterations), which promotes the progression of PC. In recent years, breakthroughs have been made in targeted therapy, including the clinical application of KRASG12C inhibitors (sotorasib, adagrasib) and KRASG12D inhibitors, and the strategies targeting epidermal growth factor receptor, DNA repair (PARP inhibitors), and immune microenvironment (combined therapies targeting PD-1 and PD-L1) have significantly improved the treatment outcome of PC. Nevertheless, drug resistance and tumor heterogeneity remain huge challenges. Precision medicine and combined therapies should be adopted in the future to improve the prognosis of patients.
-
Key words:
- Pancreatic Neoplasms /
- KRAS mutation /
- Epigenomics /
- Molecular Targeted Therapy
-
注: MAPK,丝裂原活化蛋白激酶;PI3K,磷脂酰肌醇3激酶;KRAS,Kirsten大鼠肉瘤病毒癌基因同源物;TP53,肿瘤蛋白p53;CDKN2A,细胞周期蛋白依赖性激酶抑制因子2A;SMAD4,SMAD家族成员4;PARP,多腺苷二磷酸核糖聚合酶;EGFR,表皮生长因子受体;CAF,癌症相关成纤维细胞;PRC2,多梳抑制复合物2; RNAPⅡ,RNA聚合酶Ⅱ;SAM,S-腺苷甲硫氨酸。百分数表示相应基因或蛋白发生突变或失调的概率。
图 1 胰腺癌发病机制图
Figure 1. Schematic diagram of the pathogenesis of pancreatic cancer
-
[1] ZHANG YY, YANG ZC, LIU YC, et al. Targeting lipid metabolism: Novel insights and therapeutic advances in pancreatic cancer treatment[J]. Lipids Health Dis, 2025, 24( 1): 12. DOI: 10.1186/s12944-024-02426-0. [2] SHI YY, ZHENG HL, WANG TZ, et al. Targeting KRAS: From metabolic regulation to cancer treatment[J]. Mol Cancer, 2025, 24( 1): 9. DOI: 10.1186/s12943-024-02216-3. [3] YIN GW, HUANG J, PETELA J, et al. Targeting small GTPases: Emerging grasps on previously untamable targets, pioneered by KRAS[J]. Signal Transduct Target Ther, 2023, 8( 1): 212. DOI: 10.1038/s41392-023-01441-4. [4] RECK M, CARBONE DP, GARASSINO M, et al. Targeting KRAS in non-small-cell lung cancer: Recent progress and new approaches[J]. Ann Oncol, 2021, 32( 9): 1101- 1110. DOI: 10.1016/j.annonc.2021.06.001. [5] PARIKH K, BANNA G, LIU SV, et al. Drugging KRAS: Current perspectives and state-of-art review[J]. J Hematol Oncol, 2022, 15( 1): 152. DOI: 10.1186/s13045-022-01375-4. [6] PANTSAR T. The current understanding of KRAS protein structure and dynamics[J]. Comput Struct Biotechnol J, 2020, 18: 189- 198. DOI: 10.1016/j.csbj.2019.12.004. [7] KRAMER-DRAUBERG M, PETRINI E, MIRA A, et al. Oncogenic mutant KRAS inhibition through oxidation at cysteine 118[J]. Mol Oncol, 2025, 19( 2): 311- 328. DOI: 10.1002/1878-0261.13798. [8] BURSKA AN, ILYASSOVA B, DILDABEK A, et al. Enhancing an oxidative“Trojan horse” action of vitamin C with arsenic trioxide for effective suppression of KRAS-mutant cancers: A promising path at the bedside[J]. Cells, 2022, 11( 21): 3454. DOI: 10.3390/cells1121-3454. [9] WANG C, HOU YC, HONG YK, et al. Intercellular TIMP-1-CD63 signaling directs the evolution of immune escape and metastasis in KRAS-mutated pancreatic cancer cells[J]. Mol Cancer, 2025, 24( 1): 25. DOI: 10.1186/s12943-024-02207-4. [10] KEMP SB, CHENG N, MARKOSYAN N, et al. Efficacy of a small-molecule inhibitor of KrasG12D in immunocompetent models of pancreatic cancer[J]. Cancer Discov, 2023, 13( 2): 298- 311. DOI: 10.1158/2159-8290.cd-22-1066. [11] HALLIN J, BOWCUT V, CALINISAN A, et al. Anti-tumor efficacy of a potent and selective non-covalent KRASG12D inhibitor[J]. Nat Med, 2022, 28( 10): 2171- 2182. DOI: 10.1038/s41591-022-02007-7. [12] LEE JK, SIVAKUMAR S, SCHROCK AB, et al. Comprehensive pan-cancer genomic landscape of KRAS altered cancers and real-world outcomes in solid tumors[J]. NPJ Precis Oncol, 2022, 6( 1): 91. DOI: 10.1038/s41698-022-00334-z. [13] McDAID WJ, WILSON L, ADDERLEY H, et al. The PI3K-AKT-mTOR axis persists as a therapeutic dependency in KRAS(G12D)-driven non-small cell lung cancer[J]. Mol Cancer, 2024, 23( 1): 253. DOI: 10.1186/s12943-024-02157-x. [14] MOORE AR, ROSENBERG SC, MCCORMICK F, et al. RAS-targeted therapies: Is the undruggable drugged[J]. Nat Rev Drug Discov, 2020, 19( 8): 533- 552. DOI: 10.1038/s41573-020-0068-6. [15] GRUNDY MK, BUCKANOVICH RJ, BERNSTEIN KA. Regulation and pharmacological targeting of RAD51 in cancer[J]. NAR Cancer, 2020, 2( 3): zcaa024. DOI: 10.1093/narcan/zcaa024. [16] PEREIRA F, FERREIRA A, REIS CA, et al. KRAS as a modulator of the inflammatory tumor microenvironment: Therapeutic implications[J]. Cells, 2022, 11( 3): 398. DOI: 10.3390/cells11030398. [17] ABAD E, SANDOZ J, ROMERO G, et al. The TP53-activated E3 ligase RNF144B is a tumour suppressor that prevents genomic instability[J]. J Exp Clin Cancer Res, 2024, 43( 1): 127. DOI: 10.1186/s13046-024-03045-4. [18] KENNEDY MC, LOWE SW. Mutant p53: It’s not all one and the same[J]. Cell Death Differ, 2022, 29( 5): 983- 987. DOI: 10.1038/s41418-022-00989-y. [19] TOMBARI C, ZANNINI A, BERTOLIO R, et al. Mutant p53 sustains serine-Glycine synthesis and essential amino acids intake promoting breast cancer growth[J]. Nat Commun, 2023, 14( 1): 6777. DOI: 10.1038/s41467-023-42458-1. [20] REESE KL, PANTEL K, SMIT DJ. Multibiomarker panels in liquid biopsy for early detection of pancreatic cancer-a comprehensive review[J]. J Exp Clin Cancer Res, 2024, 43( 1): 250. DOI: 10.1186/s13046-024-03166-w. [21] CAPORALI S, BUTERA A, RUZZA A, et al. Selective metabolic regulations by p53 mutant variants in pancreatic cancer[J]. J Exp Clin Cancer Res, 2024, 43( 1): 310. DOI: 10.1186/s13046-024-03232-3. [22] RAN RS, LI LX, CHENG P, et al. High frequency of melanoma in cdkn2b(-/-)/tp53(-/-) Xenopus tropicalis[J]. Theranostics, 2024, 14( 19): 7470- 7487. DOI: 10.7150/thno.97475. [23] ARAKI T, MIWA N. Selective epigenetic alterations in RNF43 in pancreatic exocrine cells from high-fat-diet-induced obese mice; implications for pancreatic cancer[J]. BMC Res Notes, 2024, 17( 1): 106. DOI: 10.1186/s13104-024-06757-0. [24] LEE HK, NAM MW, GO RE, et al. TGF-β2 antisense oligonucleotide enhances T-cell mediated anti-tumor activities by IL-2 via attenuation of fibrotic reaction in a humanized mouse model of pancreatic ductal adenocarcinoma[J]. Biomed Pharmacother, 2023, 159: 114212. DOI: 10.1016/j.biopha.2022.114212. [25] LI ZC, MA ZY, WANG SJ, et al. LINC00909 up-regulates pluripotency factors and promotes cancer stemness and metastasis in pancreatic ductal adenocarcinoma by targeting SMAD4[J]. Biol Direct, 2024, 19( 1): 24. DOI: 10.1186/s13062-024-00463-4. [26] GRAHAM S, DMITRIEVA M, VENDRAMINI-COSTA DB, et al. From precursor to cancer: Decoding the intrinsic and extrinsic pathways of pancreatic intraepithelial neoplasia progression[J]. Carcinogenesis, 2024, 45( 11): 801- 816. DOI: 10.1093/carcin/bgae064. [27] ZHAO YH, YANG M, WANG SJ, et al. An overview of epigenetic methylation in pancreatic cancer progression[J]. Front Oncol, 2022, 12: 854773. DOI: 10.3389/fonc.2022.854773. [28] LO EKW, MEARS BM, MAURER HC, et al. Comprehensive DNA methylation analysis indicates that pancreatic intraepithelial neoplasia lesions are acinar-derived and epigenetically primed for carcinogenesis[J]. Cancer Res, 2023, 83( 11): 1905- 1916. DOI: 10.1158/0008-5472.CAN-22-4052. [29] WANG SS, HALL ML, LEE E, et al. Whole-genome bisulfite sequencing identifies stage- and subtype-specific DNA methylation signatures in pancreatic cancer[J]. iScience, 2024, 27( 4): 109414. DOI: 10.1016/j.isci.2024.109414. [30] ZHOU Q, PICHLMEIER S, DENZ AM, et al. Altered histone acetylation patterns in pancreatic cancer cell lines induce subtype-specific transcriptomic and phenotypical changes[J]. Int J Oncol, 2024, 64( 3): 26. DOI: 10.3892/ijo.2024.5614. DOI: 10.3892/ijo.2024.5614. [31] WRIGHT CA, GORDON ER, COOPER SJ. Genomic analysis reveals HDAC1 regulates clinically relevant transcriptional programs in Pancreatic cancer[J]. BMC Cancer, 2023, 23( 1): 1137. DOI: 10.1186/s12885-023-11645-0. [32] LI HF, WANG HL, CUI YL, et al. EZH2 regulates pancreatic cancer cells through E2F1, GLI1, CDK3, and Mcm4[J]. Hereditas, 2023, 160( 1): 23. DOI: 10.1186/s41065-023-00280-1. [33] NEMETH K, BAYRAKTAR R, FERRACIN M, et al. Non-coding RNAs in disease: From mechanisms to therapeutics[J]. Nat Rev Genet, 2024, 25( 3): 211- 232. DOI: 10.1038/s41576-023-00662-1. [34] MOK ETY, CHITTY JL, COX TR. miRNAs in pancreatic cancer progression and metastasis[J]. Clin Exp Metastasis, 2024, 41( 3): 163- 186. DOI: 10.1007/s10585-023-10256-0. [35] WANG B, YUAN C, QIE YY, et al. Long non-coding RNAs and pancreatic cancer: A multifaceted view[J]. Biomed Pharmacother, 2023, 167: 115601. DOI: 10.1016/j.biopha.2023.115601. [36] LEFLER JE, MARELIA-BENNETT CB, THIES KA, et al. STAT3 in tumor fibroblasts promotes an immunosuppressive microenvironment in pancreatic cancer[J]. Life Sci Alliance, 2022, 5( 11): e202201460. DOI: 10.26508/lsa.202201460. [37] TOST J, AK-AKSOY S, CAMPA D, et al. Leveraging epigenetic alterations in pancreatic ductal adenocarcinoma for clinical applications[J]. Semin Cancer Biol, 2025, 109: 101- 124. DOI: 10.1016/j.semcancer.2025.01.003. [38] de LANGEN AJ, JOHNSON ML, MAZIERES J, et al. Sotorasib versus docetaxel for previously treated non-small-cell lung cancer with KRAS(G12C) mutation: A randomised, open-label, phase 3 trial[J]. Lancet, 2023, 401( 10378): 733- 746. DOI: 10.1016/S0140-6736(23)00221-0. [39] ZHANG J, JOHNSON M, BARVE M, et al. Practical guidance for the management of adverse events in patients with KRASG12C-mutated non-small cell lung cancer receiving adagrasib[J]. Oncologist, 2023, 28( 4): 287- 296. DOI: 10.1093/oncolo/oyad051. [40] TENG KW, TSAI ST, HATTORI T, et al. Selective and noncovalent targeting of RAS mutants for inhibition and degradation[J]. Nat Commun, 2021, 12( 1): 2656. DOI: 10.1038/s41467-021-22969-5. [41] AL-MAJED A. Profiles of drug substances, excipients, and related methodology[M]. Chantilly: Elsevier Science& Technology, 2023: 48. DOI: 10.1016/S1871-5125(23)00009-2. [42] LI JY, YUAN S, NORGARD RJ, et al. Epigenetic and transcriptional control of the epidermal growth factor receptor regulates the tumor immune microenvironment in pancreatic cancer[J]. Cancer Discov, 2021, 11( 3): 736- 753. DOI: 10.1158/2159-8290.CD-20-0519. [43] SHETTY SR, YEERAVALLI R, BERA T, et al. Recent advances on epidermal growth factor receptor as a molecular target for breast cancer therapeutics[J]. Anticancer Agents Med Chem, 2021, 21( 14): 1783- 1792. DOI: 10.2174/1871520621666201222143213. [44] GOLIVI Y, KUMARI S, FARRAN B, et al. Small molecular inhibitors: Therapeutic strategies for pancreatic cancer[J]. Drug Discov Today, 2024, 29( 7): 104053. DOI: 10.1016/j.drudis.2024.104053. [45] MOORE MJ, GOLDSTEIN D, HAMM J, et al. Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: A phase III trial of the national cancer institute of Canada clinical trials group[J]. J Clin Oncol, 2007, 25( 15): 1960- 1966. DOI: 10.1200/jco.2006.07.9525. [46] LEUNG PY, CHEN WJ, SARI AN, et al. Erlotinib combination with a mitochondria-targeted ubiquinone effectively suppresses pancreatic cancer cell survival[J]. World J Gastroenterol, 2024, 30( 7): 714- 727. DOI: 10.3748/wjg.v30.i7.714. [47] PHAM TD, BECKER JH, METROPULOS AE, et al. Regorafenib induces DNA damage and enhances PARP inhibitor efficacy in pancreatic ductal carcinoma[J]. BMC Cancer, 2024, 24( 1): 1562. DOI: 10.1186/s12885-024-13334-y. [48] GOLAN T, HAMMEL P, RENI M, et al. Maintenance olaparib for germline BRCA-mutated metastatic pancreatic cancer[J]. N Engl J Med, 2019, 381( 4): 317- 327. DOI: 10.1056/NEJMoa1903387. [49] LI EL, HUANG X, ZHANG G, et al. Combinational blockade of MET and PD-L1 improves pancreatic cancer immunotherapeutic efficacy[J]. J Exp Clin Cancer Res, 2021, 40( 1): 279. DOI: 10.1186/s13046-021-02055-w. [50] PAN Y, FEI QL, XIONG P, et al. Synergistic inhibition of pancreatic cancer with anti-PD-L1 and c-Myc inhibitor JQ1[J]. Oncoimmunology, 2019, 8( 5): e1581529. DOI: 10.1080/2162402X.2019.1581529. [51] LEMAITRE L, ADENIJI N, SURESH A, et al. Spatial analysis reveals targetable macrophage-mediated mechanisms of immune evasion in hepatocellular carcinoma minimal residual disease[J]. Nat Cancer, 2024, 5( 10): 1534- 1556. DOI: 10.1038/s43018-024-00828-8. [52] XIE BC, WU PS, LIU HY, et al. Long non-coding RNA MIR4435-2HG modulates pancreatic cancer stem cells and chemosensitivity to gemcitabine by targeting the miR-1252-5p/STAT1[J]. J Transl Med, 2025, 23( 1): 165. DOI: 10.1186/s12967-025-06128-8. -


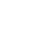 PDF下载 ( 945 KB)
PDF下载 ( 945 KB)
 下载:
下载:


