咖啡酸苯乙酯对肝星状细胞的作用及机制分析
DOI: 10.3969/j.issn.1001-5256.2022.10.014
-
摘要:
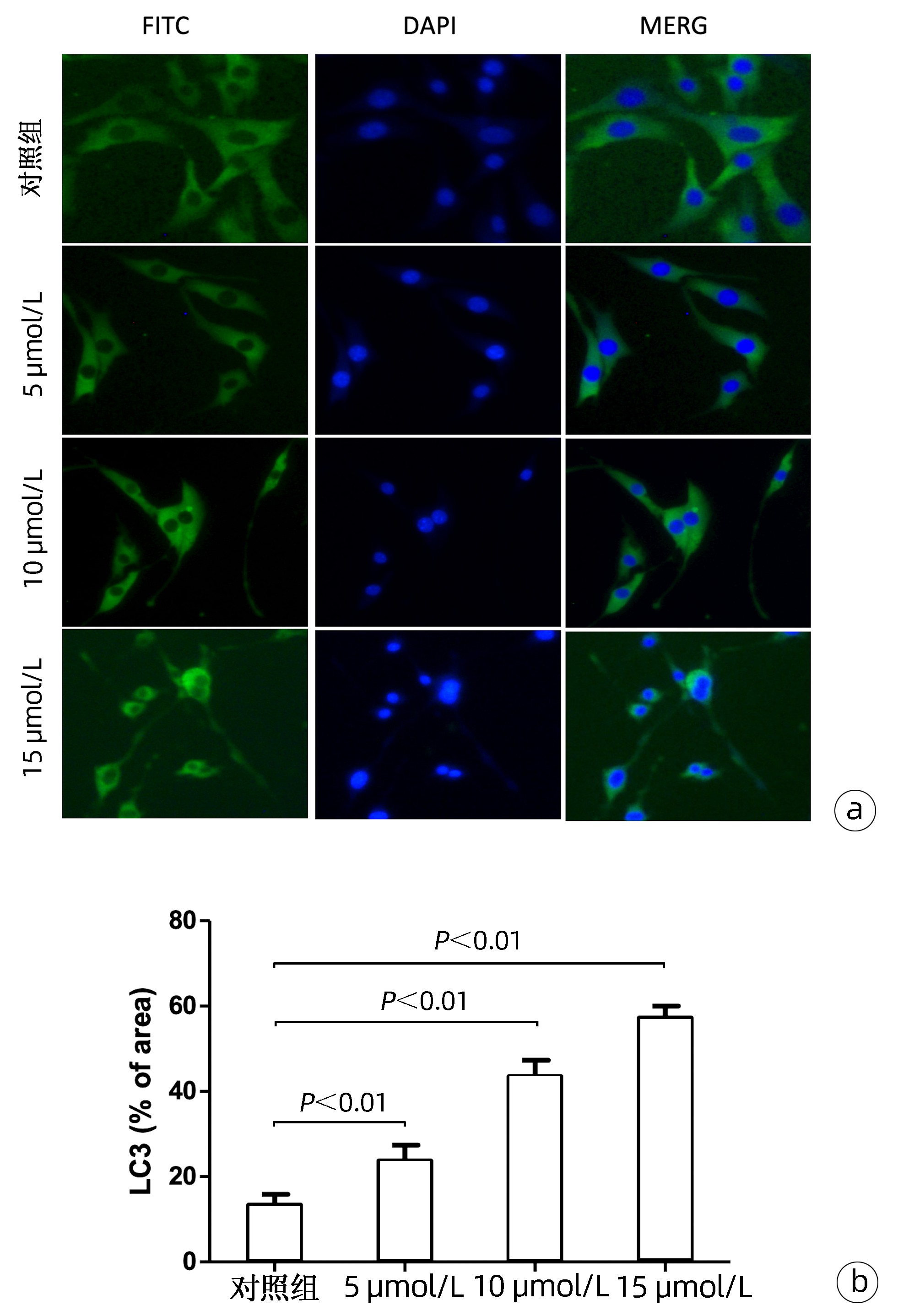
目的 探讨咖啡酸苯乙酯(CAPE)对大鼠肝星状细胞(HSC)-T6的作用及作用机制。 方法 不同浓度CAPE(5、10、15 μmol/L) 分别作用于HSC-T6细胞,同时构建GFP-LC3质粒转染至HSC-T6细胞,自噬诱导剂雷帕霉素和自噬抑制剂3-Methyladenine(3-MA)进行干预,采用MTT方法检测CAPE对HSC-T6细胞增殖的影响,透射电镜观察CAPE对HSC-T6细胞超微结构的影响,细胞免疫荧光实验检测CAPE对HSC-T6细胞LC3表达的影响,qRT-PCR检测自噬相关基因ATG5、ATG7、ATG12、Beclin1和LC3 mRNA的表达情况,Western-Blot检测ATG7、Beclin1、LC3I/Ⅱ、p-AKT/AKT、p-mTOR蛋白的表达情况。计量资料组间比较采用单因素方差分析及Dunnett-t检验。 结果 CAPE干预组较对照组可抑制HSC-T6细胞增殖,且生长明显受到抑制,细胞质中可观察到脂滴和轮盘状自噬体散在分布。与对照组(13.34%±2.59%)相比,不同浓度CAPE 5 μmol/L (23.68%±3.76%,t=-5.553,P<0.001)、10 μmol/L(43.47%±3.83%,t=-15.958,P<0.001)、15 μmol/L(57.25%±2.78%,t=-28.334,P<0.001)的LC3荧光染色均显著增强,且呈正相关。CAPE干预组(10、15 μmol/L)ATG5、ATG7、ATG12、Beclin1和LC3 mRNA水平明显增加(P值均<0.05)。GFP-LC3质粒转染至HSC-T6细胞后,CAPE干预组(79.01%±6.69%)较对照组(67.06%±6.74%)LC3荧光染色明显增强(t=-3.083,P=0.012);同时给予自噬诱导剂雷帕霉素(86.88%±5.42%),LC3表达较CAPE组明显增强(t=-2.239,P=0.049);而给予自噬抑制剂3-MA(71.22%±4.29%),其表达则明显减低(t=-2.404,P=0.037);CAPE干预组和诱导剂雷帕霉素组ATG7、Beclin1和LC3 I/Ⅱ的蛋白表达水平增加,AKT/p-AKT、p-mTOR蛋白的表达水平降低;而抑制剂3-MA组则得出相反的结果。 结论 CAPE可能通过抑制AKT/mTOR信号通路诱导自噬形成,进而抑制HSC-T6细胞增殖。 Abstract:Objective To assess the effect and underlying molecular events of caffeic acid phenethyl ester (CAPE) on rat hepatic stellate HSC-T6 cells. Methods HSC-T6 cells were grown and treated with different concentrations of CAPE (5, 10, or 15 μmol/L), transfected with or without LC3-GFP plasmid, and then treated with or without an autophagy inducer rapamycin or the autophagy inhibitor 3-methyladenine (3-MA). The changed cell viability and morphology were assessed by using cell viability MTT assay and Transmission electron microscope, respectively. The expression of LC3 protein in HSC-T6 cells was detected by immunofluorescence assay, the autophagy-related genes expression of ATG5, ATG7, ATG12, Beclin1 and LC3 were detected by qRT-PCR, and the expression of ATG7, Beclin1, LC3I/Ⅱ, p-AKT/AKT, p-mTOR protein was detected by Western-blot. Comparison between multiple groups was analyzed by one-way ANOVA with Dunnett t-test. Results Compared with the control, CAPE treatment significantly reduced cell viability but induced formation of lipid droplets and roulette-shaped autophagosomes. Compared with the control (13.34%±2.59), LC3 protein was significantly induced in HSC-T6 cells after CAPE treatment (5 μmol/L, 23.68%±3.76, t=-5.553, P < 0.001; 10 μmol/L, 43.47%±3.83, t=-15.958, P < 0.001; 15 μM, 57.25%±2.78, t=-28.334, P < 0.001), while levels of ATG5, ATG7, ATG12, Beclin 1, and LC3 mRNAs were all significantly increased in 10 μm and 15 μm CAPE treated cells vs the control (all P < 0.05). After LC3 overexpression in HSC-T6 cells, LC3 protein was induced vs the vector control (79.01%±6.69% vs 67.06%±6.74%, t=-3.083, P=0.012), while rapamycin treatment further increased LC3 expression (86.88%±5.42%, t=-2.239, P=0.049); however, 3-MA treatment significantly decreased LC3 expression in cells (71.22%±4.29%, t=-2.404, P=0.037). In addition, levels of ATG7, Beclin1, and LC3 Ⅰ/Ⅱ proteins were increased, whereas levels of AKT/p-AKT and p-mTOR were decreased in the CAPE and rapamycin groups vs controls. However, the 3-MA treatment had an opposite result, indicating that 3-MA reversed CAPE-induced effects in HSC-T6 cells. Conclusion Caffeic acid phenethyl ester may induce autophagy to reduce cell viability in hepatic stellate cells by inhibition of the AKT/mTOR signaling. -
Key words:
- Liver Cirrhosis /
- Caffeic Acids /
- Hepatic Stellate Cell
-
表 1 Real-time PCR引物序列
Table 1. Primes of Real-time PCR
基因 引物 温度(℃) 长度(bp) 来源 ATG5 F 5′-TCAGTGGAGGCAACAGAACC-3′ 55.49 20 大鼠 R 5′-TTCACTGAGCAAAAGCGTGC-3′ 55.71 20 大鼠 ATG7 F 5′-GAGACCGTCTGAGCAACCAC-3′ 56.38 20 大鼠 R 5′-GATGACACAGGAAAGGGTGC-3′ 54.5 20 大鼠 ATG12 F 5′-CGGACTGTCCAAGCACTCAT-3′ 55.68 20 大鼠 R 5′-CCCATGCCTGTGATTTGCAG-3′ 55.49 20 大鼠 LC3 F 5′-CGGGTTGAGGAGACACACAA -3′ 55.53 20 大鼠 R 5′-ATGAGCCGGACATCTTCCAC -3′ 55.44 20 大鼠 Beclin1 F 5′-GAATGGAGGGGTCTAAGGCG -3′ 55.52 20 大鼠 R 5′-GAATGGTCACTCGGTCCAGG -3′ 55.76 20 大鼠 β-actin F 5′-GGAGATTACTGCCCTGGCTCCTA-3′ 60.2 23 大鼠 R 5′-GACTCATCGTACTCCTGCTTGCTG-3′ 59.3 24 大鼠 表 2 CAPE对自噬相关的基因表达的影响
Table 2. The expression levels of autophagy-related genes in HSC-T6 cells with CAPE treatment
指标 对照组 CAPE F值 方差分析的P值 5 μmol/L 10 μmol/L 15 μmol/L ATG5 1.02±0.01 1.06±0.04 1.46±0.091) 2.43±0.421) 15.770 <0.05 ATG7 1.05±0.02 1.08±0.04 3.30±0.231) 5.44±0.151) 665.826 <0.01 ATG12 1.01±0.06 1.07±0.03 2.33±0.081) 3.80±0.171) 585.839 <0.01 Becline1 0.99±0.08 1.07±0.05 1.55±0.071) 2.44±0.131) 182.940 <0.01 LC3 1.08±0.04 1.12±0.10 1.55±0.111) 2.38±0.271) 44.963 <0.01 注:与对照组比较,1)P<0.01。 -
[1] MURTAZA G, KARIM S, AKRAM MR, et al. Caffeic acid phenethyl ester and therapeutic potentials[J]. Biomed Res Int, 2014, 2014: 145342. DOI: 10.1155/2014/145342. [2] OLGIERD B, KAMILA Z, ANNA B, et al. The pluripotent activities of caffeic acid phenethyl ester[J]. Molecules, 2021, 26(5): 1335. DOI: 10.3390/molecules26051335. [3] LI M, WANG XF, SHI JJ, et al. Caffeic acid phenethyl ester inhibits liver fibrosis in rats[J]. World J Gastroenterol, 2015, 21(13): 3893-3903. DOI: 10.3748/wjg.v21.i13.3893. [4] YANG N, SHI JJ, WU FP, et al. Caffeic acid phenethyl ester up-regulates antioxidant levels in hepatic stellate cell line T6 via an Nrf2-mediated mitogen activated protein kinases pathway[J]. World J Gastroenterol, 2017, 23(7): 1203-1214. DOI: 10.3748/wjg.v23.i7.1203. [5] HAZARI Y, BRAVO-SAN PEDRO JM, HETZ C, et al. Autophagy in hepatic adaptation to stress[J]. J Hepatol, 2020, 72(1): 183-196. DOI: 10.1016/j.jhep.2019.08.026. [6] KOUROUMALIS E, VOUMVOURAKI A, AUGOUSTAKI A, et al. Autophagy in liver diseases[J]. World J Hepatol, 2021, 13(1): 6-65. DOI: 10.4254/wjh.v13.i1.6. [7] ALLAIRE M, RAUTOU PE, CODOGNO P, et al. Autophagy in liver diseases: Time for translation?[J]. J Hepatol, 2019, 70(5): 985-998. DOI: 10.1016/j.jhep.2019.01.026. [8] ROEHLEN N, CROUCHET E, BAUMERT TF. Liver fibrosis: mechanistic concepts and therapeutic perspectives[J]. Cells, 2020, 9(4): 875. DOI: 10.3390/cells9040875. [9] KISSELEVA T, BRENNER D. Molecular and cellular mechanisms of liver fibrosis and its regression[J]. Nat Rev Gastroenterol Hepatol, 2021, 18(3): 151-166. DOI: 10.1038/s41575-020-00372-7. [10] LEVINE B, KROEMER G. Biological functions of autophagy genes: a disease perspective[J]. Cell, 2019, 176(1-2): 11-42. DOI: 10.1016/j.cell.2018.09.048. [11] WANG H, LIU Y, WANG D, et al. The upstream pathway of mTOR-mediated autophagy in liver diseases[J]. Cells, 2019, 8(12): 1597. DOI: 10.3390/cells8121597. [12] LEE KW, THIYAGARAJAN V, SIE HW, et al. Synergistic effect of natural compounds on the fatty acid-induced autophagy of activated hepatic stellate cells[J]. J Nutr Biochem, 2014, 25(9): 903-913. DOI: 10.1016/j.jnutbio.2014.04.001. [13] PARK M, KIM YH, WOO SY, et al. Tonsil-derived mesenchymal stem cells ameliorate CCl4-induced liver fibrosis in mice via autophagy activation[J]. Sci Rep, 2015, 5: 8616. DOI: 10.1038/srep08616. -


 PDF下载 ( 3612 KB)
PDF下载 ( 3612 KB)

 下载:
下载: